International Journal of Veterinary Science and Research
The identification of gene ontologies and candidate genes for digital dermatitis in beef cattle from a genome-wide association study
1Institute of Agricultural and Nutritional Sciences, University of Halle, Halle, Germany, 06099
2Department of Medical Science, School of Veterinary Medicine, University of Wisconsin, Madison, WI 53706, USA
3Department of Animal Sciences, University of Wisconsin, Madison, WI 53706, USA
Author and article information
Cite this as
Kopke G, Anklam K, Kulow M, Baker L, Swalve HH, et al. (2020) The identification of gene ontologies and candidate genes for digital dermatitis in beef cattle from a genome-wide association study. Int J Vet Sci Res. 2020; 6(1): 27-37. Available from: 10.17352/ijvsr.000050
Copyright License
© 2020 Kopke G, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Bovine Digital Dermatitis (DD) is an infectious disease causing severe lameness in cattle. The aim of this study was to perform a Genome Wide Association Study (GWAS) and a Gene-Set Enrichment Analysis (GSEA) to identify candidate genes, instead of an individual Single Nucleotide Polymorphism (SNP), associated with DD traits in beef cattle. Beef cattle (n= 307) were genotyped with the Illumina GGP-HD bovine 150K SNP chip. The M-scores of the cattle over the observation period were used to define the DD traits with different complexities, the distinction between affected (1) and unaffected (0) cattle, regarding the general DD-status (DD AFFECTED), acute disease events (DD ACUTE), visible signs of chronicity (DD CHRONICITY) and proliferation of the skin (DD PROLIFERATION). The gene-set enrichment analysis revealed 30 Gene Ontology (GO) terms associated with the DD AFFECTED trait and 17, 31, and 16 GO terms were associated with DD ACUTE, DD CHRONICITY, and DD PROLIFERATION traits, respectively. By searching the significantly enriched GO terms from the ontology categories, biological process and cellular components, and molecular function, 25 functional genes were identified that were highly involved in cellular and membrane function pertaining to adhesion, migration and proliferation which could contribute to DD traits. These results could provide insight into the genetic framework of this complex trait and disease in beef cattle to aid the development of potential genetic therapies as well as selective breeding strategies to decrease DD prevalence in cattle.
Bovine Digital Dermatitis (DD) is a worldwide infectious disease that causes severe lameness in cattle across all production systems [1-4]. The consequences of DD are decreased animal welfare and economic losses due to reduced milk production, decreased reproductive performance and premature culling [5,6]. DD has a multifactorial etiology with the primary causative infectious agent being Treponema spp. and the pathogenesis of DD is not completely understood [7,8]. DD occurs in different clinical manifestations and recurrence after therapy is high, resulting in problems for prevention and control [1,9]. Moderate to high heritability estimates for DD traits based on the M-stage system and varying immune responses supports the influence of host genetic factors [10-14]. Genome wide association and gene expression studies have been performed to identify potential candidate genes with influence on DD, however causative mutations have yet to be defined based on the current literature [15-17]. Gene-set enrichment and pathway-based analyses have been described to investigate the polygenic background of complex traits, such as leucosis, bull fertility, and beef cattle meat quality [18-20]. DAVID 6.8 is a web based tool to discover enriched functionally related gene groups for large gene lists and for their interpretation in biological context [21]. Gene-set enrichment directs the focus from a single gene-oriented view to a gene-group-based analysis. Findings of these analyses can contribute to an improved understanding about the genetic and biological architecture of complex traits, such as DD. In addition, these findings could be useful for future genetic prevention strategies against DD and an improved management of cows susceptible to DD would benefit from gene-set enrichment analysis. The objective of this study was to perform a Genome Wide Association Study (GWAS) for DD in beef cattle and to identify candidate genes using a gene-set enrichment based analysis.
Material and methods
The experimental procedures for the trial were approved by the Institutional Animal Care and Use Committee (IACUC V01525) at the University of Wisconsin-Madison.
Study design
Data were collected between June 2014 and November 2015 during randomized field trials to evaluate the effect of an Organic Trace Mineral (OTM) (Availa Plus; Zinpro Performance Minerals, Eden Prairie, MN) program on the prevalence of DD in two commercial feedlot farms (Midwest, US) [22]. The producers had observed increasing numbers of outbreaks of DD over the past five years on these farms. The highest occurrence of DD outbreaks were observed during the summer months 60 days prior to slaughter. Cattle were housed in barns covered by monoslopes on concrete, grooved flooring and a bedded pack in the center. Cattle were randomly split over 19 pens, 7 control pens receiving a diet supplemented with trace minerals of inorganic sources and 12 OTM pens receiving the same diet supplemented with organic trace minerals (OTM). A total of 1,120 heifers (farm A) and 1,077 steers (farm B) were enrolled in the study. Cattle were mixed breeds, and were assigned into a “breed” category based on coat color and were sourced from multiple locations in North America. Pens of cattle selected to be in the study were processed for enrollment using a restraint chute (Silencer, Moly Manufacturing Inc., Lorraine, KS). Individual weights were recorded for all cattle enrolled and tail hair follicle samples were taken from 2,197 animals for genotyping.
While cattle were in the chute, their hind feet were evaluated for the presence of DD and scored based on DD lesion type using the M-stage DD classification system described by Döpfer and Berry [23,24]. Based on this classification system cattle with: normal skin appearance observed on the foot were classified as M0; active ulcerative DD lesions ≥ 20 mm in diameter were classified as M2 lesions and chronic lesions were classified as M4. Further signs of chronicity were classified as: none = smooth skin without thickening and no extra tissue beyond the surface of the normal level of the skin; hyperkeratotic = thickened callous skin; and proliferative = diffuse, filamentous or mass-like overly grown epidermal tissues. If both hind legs were affected, or more than one lesion was present during scoring, the more severe lesion was documented (M2 > M4 > M0 in severity, proliferative lesions were considered more severe than hyperkeratotic lesions). The previously described DD Check App was used for data documentation [25]. Collection of data was conducted by two trained investigators. All hind feet of cattle were evaluated during single scoring events, three in the chute and three during alley checks, for which where groups of 3 to 5 cattle were herded into an alleyway.
Phenotype and genotype data
The phenotype data set contained 9,653 observations of 2,197 cattle from farms A and B. The cattle that had at least two observations for DD-status with the majority of cattle (85.3%) having four and more observations were used for analysis. Table 1 shows an overview of characteristics and average prevalence of M-Stages and signs of chronicity per farm. Binary traits were defined for the distinction between affected (1) and unaffected (0) cattle, regarding the general DD-status (DD AFFECTED), acute disease events (DD ACUTE), visible signs of chronicity (DD CHRONICITY) and proliferation of the skin (DD PROLIFERATION). Table 2 shows an overview of trait definitions and overall DD prevalence.
DNA was extracted from approximately 30 hair follicles from each animal using the DNeasy blood and tissue kit (Qiagen, Valencia, CA) according to the manufacturer’s instructions. Although hair follicle samples for genotyping were collected from 2,197 animals unfortunately only 307 samples yielded the required concentration of DNA for analysis. The genotype analysis was performed after the completion of the study therefore we were unable to recollect the samples. Genotyping was performed on the DNA by GeneSeek (Neogen Genomics, Lincoln, NE) utilizing the GGP-HD Bovine Illumina 150K Chip a genome-wide genotyping array. The genotype data were edited using PLINK v. 1.07 [26]. Single nucleotide polymorphisms (SNPs) with a minor allele frequency < 0.05, not in Hardy Weinberg equilibrium (p < 0.0000001) and with missing call rates exceeding 5% were removed. As the data contained both steers and heifers, the X chromosome was excluded from further analysis. A total of 117,479 SNP on Bos taurus autosome (BTA) 1 to 29 were available for final analysis. All the missing SNP information on these SNPs (0.27%) was imputed using FImpute in R [27].
Statistical analysis and modeling
Computation of a genomic relationship matrix (G-matrix) as well as a principal component analysis (PCA) and a cluster analysis using a Bray Curtis distance matrix and UPGMA clustering were performed with the packages “rrBLUP” and “vegan” in R version 3.3.3 [28-30].
Model testing was performed for the fixed effects (farm, coat color, treatment, and cluster) and covariates (3 principal components and initial body weight) as well as their interactions. The simplest model regarding fixed effects with the lowest Akaike information criterion (AIC) was chosen:
logit(pijkl) = μ + farmi + coatclj + treatmentk + β1(InitialWtl) + β2(PC1l) + β3(PC2l) + β4(PC3l) (1)
where pijkl = P(Yijkl = 1) is the probability of occurrence, Yijkl is the DD-trait (DD AFFECTED, DD ACUTE, DD CHRONICITY, DD PROLIFERATION), μ is the overall mean, farmi describes the fixed effect of farm i(i = A or B); coatclj is the fixed effect of coat color j (j = black or others), treatmentk is the fixed effect of treatment k (k = CON or OTM), β1(InitialWt1) is the regression coefficient for initial body weight of animal l and β2(PC1l), β3(PC2l), β4(PC3l) are the regression coefficients for the first three principal components from PCA of all SNPs. Least Square Estimates were calculated for initial body weight as well as Least Square Means (LSMeans) and odds ratios (OR) for farm, treatment, and coat color using a logit link function in the “GLIMMIX” Procedure in SAS version 9.4 [31].
Genome-wide association study, identification of candidate genes and gene-set enrichment analysis
A GWAS was performed for each binary trait using the “easyGWAS” [32] package in R version 3.3.3 applying the following model:
y = μ+Wv+Xβi+Zu+e,
Where μ is the population mean, v is the vector of fixed effects, βi is the effect of the ith SNP, u is the vector of the polygenic effects, e is the residual, W, X, and Z are the incidence matrices for v, βi, and u. SNPs with a p-value < 0.01 were assumed to be statistically significant and a list of these SNPs was generated for every single trait. For enrichment analysis statistically significant SNPs were assigned to genes within 20kb upstream and downstream of the gene’s start and end position as well as within the coding region of the gene using the “biomaRt” Package in R version 3.3.3 [33,34]. The distance of 20kb was chosen to include proximal functional and regulatory regions in the analysis. If a SNP was assigned to more than one gene, all assigned genes remained for further analysis. The Bos taurus UMD3.1: Release 90 - August 2017 from Ensembl was used as reference genome [35,36]. For enrichment analysis a list of the genes identified by statistically significant SNPs was submitted to DAVID 6.8 for each trait separately [21,37]. The Bos taurus gene list of DAVID 6.8 was used as the reference genome. To determine functional categories the Gene Ontology (GO) database was used [38,39]. The GO database offers biological descriptors (GO terms) for genes based on the characteristics of encoded proteins. The terms are assigned to three categories: biological process (BP), Molecular Function (MF) and Cellular Component (CC) [38,40]. A Fisher’s exact test was performed to search for an overrepresentation of significantly associated genes among all the genes in the given GO term. Finally only functional enriched terms with a Fisher’s exact test <0.001 were considered statistically significant and used for further interpretation. GO Plots were created in R for all binary traits using the “GO:” Database and the documented ancestors of the GO terms to show connections between the functional categories. Description of single GO terms was provided by the database AmiGO 2 version 2.4.26 [41]. For identification of candidate genes the lists of statistically significant genes per trait from enrichment analysis were used to perform an extensive literature search in the National Center for Biotechnology Information (NCBI) database for the following gene involvement terms: inflammation, adhesion, skin, keratinocyte, wound healing, hyperkeratosis, proliferation, and immune response.
Results and discussion
Statistical analysis
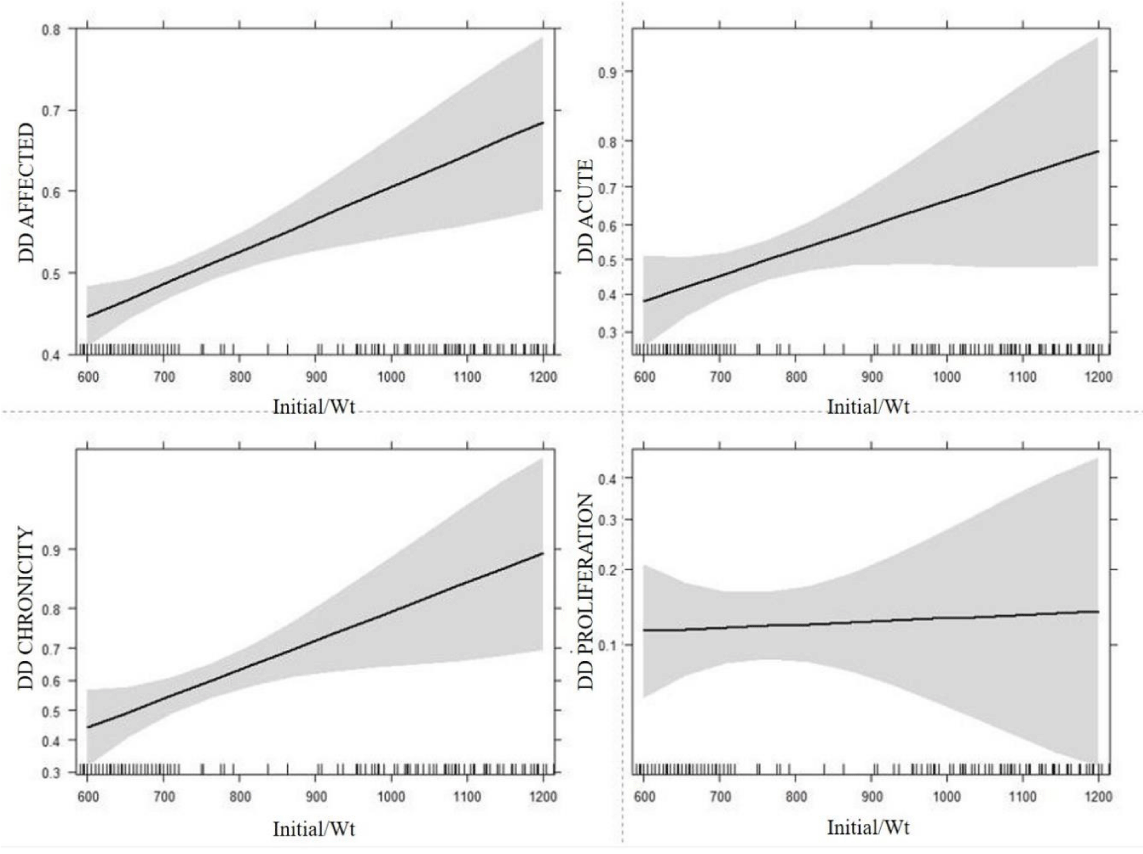
Estimates and confidence intervals for the fixed effects of farm, coat color and OTM treatment for the binary traits are presented in Table 3. Higher values were obtained for farm A as well as for Treatment CON for all traits except DD PROLIFERATION, with the higher values being statistically significant for farm A and the traits DD AFFECTED and DD CHRONICITY. For coat color, estimates were found for black coated animals in DD CHRONICITY and DD PROLIFERATION. The Least Square Estimate for the regression on initial body weight for the probability of the binary traits is presented in Figure 1. As expected, all binary traits showed a significant increase in probability as initial body weight increased, except for DD PROLIFERATION for which only a slight increase of probability was observed. These associations are indicated by the positive slope in each binary trait panel of Figure 1, while the shaded area depicts the 95% confidence interval of the estimated slope. Significant odds ratios (OR) were calculated for the influence of farm as well as a gain in 200 lbs./91 kg in initial body weight for the trait DD AFFECTED (Table 4). In addition, Table 4 shows the significantly different OR calculated for the influence of all fixed effects and initial body weight on DD CHRONICITY and coat color on DD PROLIFERATION.
As can be seen in Table 4, the farm had the greatest influence on the occurrence of DD in this study. Farm differences are understandable, because DD has a multifactorial etiology, highly influenced by management and hygiene [42]. Furthermore the results indicate that coated color was associated with the occurrence of DD, with a higher prevalence of chronic and proliferative lesions in black coated cattle (see coat color vs. DD PROLIFERATION in Table 4). If we consider coat color as a proxy for breed, this agrees with the influence of breed on DD, which has been previously described [42-44]. Several other studies described the importance of animal genetic factors and breed on DD as well [12,45]. In the current study it was not possible to determine the exact breed because no pedigree information was available.
PCA and cluster analysis
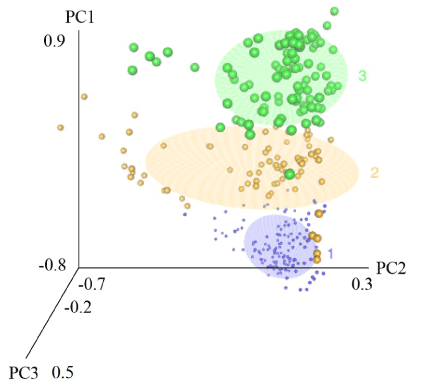
Principal Component (PC) analysis and cluster analysis resulted in three principal components and three clusters based on the genomic relationship matrix. The first three principal components explained 26.5% of the genomic variation in the data set. Figure 2 illustrates a 3D plot for the site scores of the first three principal components containing the three previously generated clusters. Clusters 1, 2, and 3 are shown as purple, yellow, and green clusters of different sized data points where the size of the data points simply represents the different clusters. The 3D plot indicates that the three clusters are mostly determined by PC1. Since the AIC of GWAS model was lowest when including the three principal components from Figure 2, those three principal components were chosen as predictors for the final GWAS model, not the cluster numbers.
Genome-wide association study and gene mapping
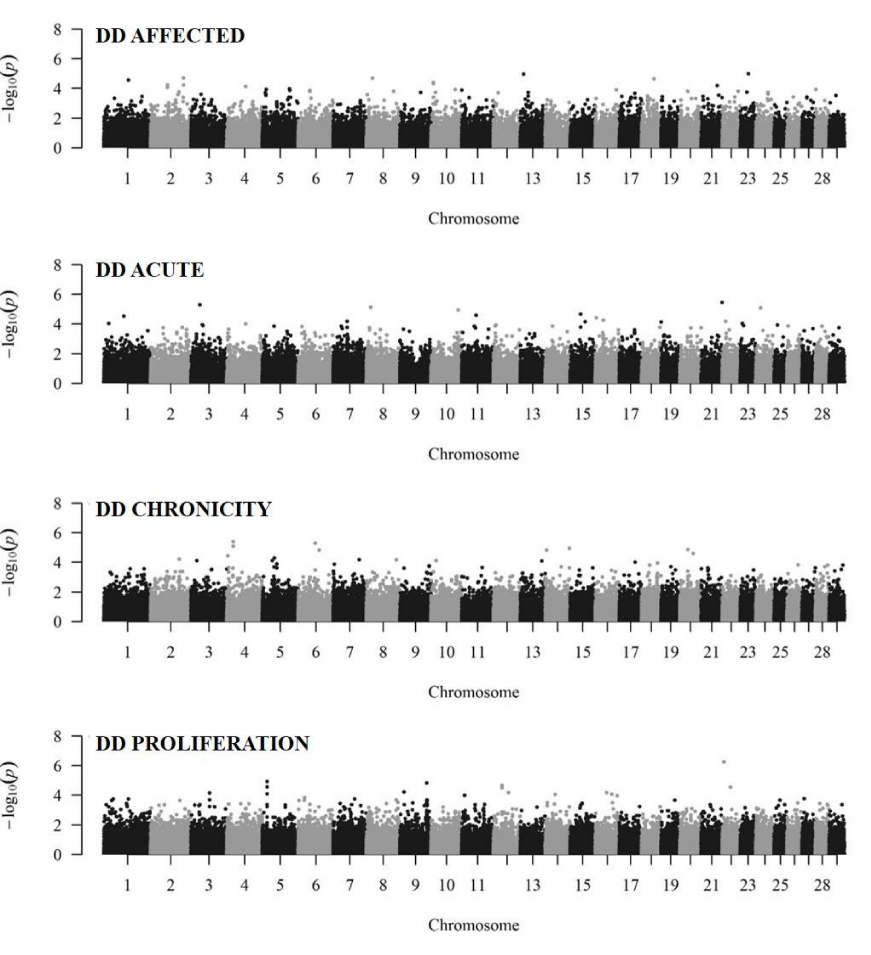
Manhattan plots illustrate the genome wide association for the binary traits in Figure 3. The genome wide suggestive level was calculated to be 1.7×10-6 corresponding to a Bonferroni-adjusted p-value of 0.2. Only one SNP (rs110651789, BTA22:5833750) showed a genome wide suggestive association to the trait DD PROLIFERATION (p-value < 1.7×10-6). No candidate genes were identified in a 20kb window surrounding this SNP. The low number of associated SNPs was expected due to the limited sample size. Therefore, for the gene set enrichment analysis the p-value of < 0.01 was used as significance criterion to ensure the necessary required number of significant SNPs for the ongoing analysis, as it was stated that gene set enrichment should include 100 to 2000 genes [37]. Table 5 shows the number of statistically significant SNPs and the number of assigned significant genes per binary trait based on the p-value (< 0.01). An SNP can be assigned to more than one gene therefore the number of significant SNPs assigned to genes is higher than the number of (unique) significant mapped genes. The 708 - 849 unique significant genes (depending on the trait) could be used for enrichment and pathway based analysis. Unfortunately, not all of these unique significant genes could be mapped in DAVID 6.8 resulting in a reduced number of significant mapped genes. For all binary traits 2,637 genes were identified to be within 20kb of a significant SNP. Due to the limited sample size we did not include a correction for multiple testing in this analysis and the results were analyzed based on the raw p-values to maximize the data available from this study. Although the high number of expected false positive SNPs from the GWAS without correction for multiple testing reduces the strength of the analysis, this is the first described genomic study for DD in a beef cattle data set. We consider our work to be an example for follow-up studies about the genetics of DD in beef cattle.
Candidate genes and enrichment analysis
A Fisher’s exact test was performed to search for an overrepresentation of significantly associated genes within each GO term. Only functional enriched terms with a Fisher’s exact test <0.001 were considered significant and used for further interpretation. A total of 13,824 background genes were assigned to the Biological Process (BP) GO term category, 15,461 background genes to the Cellular Component (CC) GO term category, and 13,094 background genes to the Molecular Function (MF) GO term category. All background genes were tested for significant enrichment of genes connected with DD in beef cattle. For the binary trait DD AFFECTED 478, 526, and 444 genes from the list of significant genes were mapped to the BP, CC, and MF gene ontology categories respectively. Slightly different numbers of genes were mapped to the BP, CC, and MF gene ontology categories for the other binary traits, DD ACUTE (580, 589, 503), DD CHRONICITY (472, 520, 431), and DD PROLIFERATION (431, 463, 380) (Online Resources 1, 2, 3). To include all possibly involved ontologies we decided to not exclude rare and widely distributed GO terms from our analysis. This is in agreement with previously published reports about GSEA in cattle [46]. Although these categories seem to be functionally broad and encompass up to 4,149 genes of the Bos taurus genome, genes within 20kb distance to a significant SNP and with functions in relation to cows individual DD-status are found within these categories. From the significantly enriched genes 25 highly relevant genes were identified by their function described in literature. Their relations to the significant SNP from GWAS and identified significantly enriched gene ontologies are provided in Table 6. The genes will be numbered 1 to 25 in the following text. For each gene ontology category (BP, CC, and MF) we will list the statistically significant enriched GO terms for the binary traits and describe associated genes from literature followed by briefly discussing their general link to the pathogenesis of DD.
GO terms significantly enriched for the category biological process (BP)
In the category BP 21 GO terms showed significant overrepresentation of genes associated with general DD-status (DD AFFECTED). Furthermore 8, 17, and 6 GO terms were significantly enriched for the binary traits DD ACUTE, DD CHRONICITY, and DD PROLIFERATION, respectively (Online Resource 1).
GO term group – localization and transport
The term localization (GO: 0051179) is significantly (Fisher’s Exact < 0.001) enriched for three different traits (DD AFFECTED, DD ACUTE, and DD CHRONICITY). Two subtypes of this term are significantly enriched in this analysis for the trait DD AFFECTED: single-organism localization (GO: 1902578) and establishment of localization (GO: 0051234). These terms are connected to localization of cells, substances, or cellular entity (protein complex or organelle) and show a close hierarchy with ontologies related to transport (GO: 0006810, GO: 0044765). The transport function is the directed movement of substances or cellular components within a cell, or between cells, or within a multicellular organism by a transporter, pore or motor protein. Cellular localization and transport play a critically important role in regulating cellular interactions. Five candidate genes were represented in this biological process and may play a role in DD pathogenesis: (1) ATG4C, (2) DDR1, (3) DOCK1, (4) ,LASP1, and (5) LIMD1. Table 7 lists the function of these five genes.
GO term group – cell differentiation
Ontologies related to the regulation of differentiation of adipose cells (GO: 0090335), GO: 0045598) and the differentiation hair cells (GO: 0035315) were significantly enriched for the trait DD AFFECTED. Further negative regulation of differentiation of keratinocytes (GO: 0045617) showed significant enrichment for the trait DD PROLIFERATION as well as a close hierarchical connection to the two previously mentioned differentiation terms. Ancestors of this term show significant enrichment for the trait DD CHRONICITY including developmental process (GO: 0032502) as well as the development of anatomical structure (GO:0048856), and nervous system (GO: 0007399). Keratinocyte differentiation is crucial for the formation of the epidermal barrier to the environment. Disturbances of the epidermal barrier formation and the dysregulation of keratinocyte differentiation are involved in the pathophysiology of several skin diseases including DD [8]. The following nine genes are involved in keratinocyte or myofibroblast differentiation: (6) DSC3, (7) CSTA, (8) COL11A1, (9) COL13A1, (10) TRAM2, (11) PRTN3, (12) SRSF6, (13) PDGFA, and (14) EGFR (Table 7). Any dysfunction in these genes could lead to an impaired epidermal barrier and allow for signs of DD to occur and to persist [24].
GO terms significantly enriched for the category cellular components (CC)
For the category CC, four GO terms showed significant overrepresentation of genes associated with general DD-status (DD AFFECTED). In addition 2, 4, and 1 GO terms were significantly enriched for the traits DD ACUTE, DD CHRONICITY, and DD PROLIFERATION, respectively (Online Resource 2).
GO term group – plasma membrane
The term integral component of plasma membrane GO: 0005887 was significantly enriched for the DD AFFECTED trait and is a subtype of GO: 0031226 intrinsic component of plasma membrane which was significantly enriched for the traits DD AFFECTED and DD CHRONICITY. GO: 0031226 is a subtype of GO: 0044459 plasma membrane part which is also significantly enriched for the DD AFFECTED trait. These terms are in close hierarchical relationship to the term GO: 0031225 anchored component of membrane which is significantly enriched for the trait DD CHRONICITY.
Plasma membrane receptors are essential for signaling many important biological events, including cell migration [47]. Motility requires adhesive interactions of cells with the extracellular matrix (ECM). These interactions are partly mediated by integrins, a family of cell surface adhesion receptors. Integrin-mediated cell adhesion regulates a multitude of cellular responses, including proliferation, survival, and cross-talk between different cellular pathways. Integrins confer different cell adhesive properties, particularly with respect to focal adhesion formation and motility [48]. Six genes are represented in the component plasma membrane term of the category cellular component: (15) ITGA1, (16) ITGA2, (17) ITGB5, (18) ITGA11, (19) ITGAV, and (20) PTGER4 (Table 7). Theses candidate genes are involve in integrin-mediated cell adhesion and cell migration and could affect DD prognosis and pathogenesis.
GO terms significantly enriched for the category molecular function (MF)
With regards to the category MF, five GO terms showed significant overrepresentation of genes associated with general DD-status (DD AFFECTED). Furthermore 7, 10, and 9 GO terms were significantly enriched for the traits DD ACUTE, DD CHRONICITY, and DD PROLIFERATION, respectively (Online Resource 3).
GO term group – transmembrane transport/channel
Ontologies related to transmembrane transport activity (GO:0022857) included the following subtypes: metal ion (GO: 0046873), inorganic cation (GO: 0022890), cation (GO: 0008324), ion (GO: 0015075), and substrate-specific (GO: 0022891 and GO: 0022892) and the ontologies were significantly enriched for the traits DD CHRONICITY and partly for DD AFFECTED.
Transmembrane transport plays a pivotal role in the communication between cells. Cells must be able to communicate across their membrane barriers to exchange materials with the environment [49]. Calcium dynamics play a key role in keratinocyte differentiation and epidermal homeostasis [50]. Impairment of these genes could interrupt epidermal homeostasis allowing for the more rapid progression of DD. The following relevant genes were included in the term category molecular function and associated with transmembrane transport/channel, (21) RYR1 and (22) TRPV4 (Table 7).
GO term group – cellular enzyme and receptor activity
The term transmembrane-ephrin receptor activity GO: 0005005 was significantly enriched for the trait DD PROLIFERATION and is related to protein kinase activity GO: 0004672 which was significantly enriched for the trait DD AFFECTED and DD PROLIFERATION. The ontologies related to transferase activity (GO: 0016740 and GO: 0016772) and the following subtypes: phosphotransferase activity (GO: 0016773) kinase activity (GO: 0016301) were also significantly enriched for the trait DD PROLIFERATION. The previously mentioned terms are in close hierarchical relationship to phosphoric ester hydrolase activity (GO: 0042578) and hydrolase activity (GO: 0016788) which were significantly enriched for the traits DD AFFECTED and DD ACUTE. Furthermore, the term collagen receptor activity (GO: 0038064) that was significantly enriched for the trait DD AFFECTED also showed close relationship to the previously mentioned terms.
Epidermal development, homeostasis, and repair are complex processes requiring regulation of cell proliferation, migration, and differentiation. Several of the signals essential for epidermal development and repair are tranduced by the epidermal growth factor receptor (EGFR) and integrins [51]. Two genes were represented in the cellular enzyme and receptor activity term of the category molecular function, (23) RAF1 and (24) ROCK1 (Table 7). These genes could influence the barrier function and integrity of the skin as well as proliferation of the skin in chronic DD lesions.
GO term group – enzyme regulator activity
Ontologies related to the terms GTPase regulator activity GO: 0030695, nucleoside-triphosphatase regulator activity GO: 0060589, and protein kinase activator activity GO: 0030295 which are subtypes of molecular function regulator GO: 0098772 were found significantly enriched for the traits DD ACUTE, DD CHRONICITY, and DD PROLIFERATION. The RAS pathway is essential to epidermal homeostasis. Epidermal function requires the balance between keratinocyte proliferation, adhesion to ECM proteins and terminal differentiation be tightly regulated. This process of balancing proliferation, adhesion, and differentiation is profoundly disrupted during pathogenesis of DD. One gene was represented in the enzyme regulator activity term of the category molecular function, (25) DAB2IP (Table 7).
The incidence of DD is a complex trait controlled by multiple genes. Therefore this approach of searching the significantly enriched GO terms and identifying 25 functional genes highly involved in cellular and membrane function pertaining to adhesion, migration and proliferation which could contribute to DD traits provides a unique opportunity to better understand the genetic architecture of this complex disease. Further pathway analysis research is needed to explore the associated biochemical, molecular, and cellular pathways to link the predominant pathways with the 25 candidate genes. This could reveal factors that contribute simultaneously to the complex trait of DD.
Conclusion
In this study a genomic analysis was performed for DD incidence in beef cattle. Digital dermatitis data were analyzed combining phenotype and genetic marker information into genomic models for the assessment of genetic parameters and for the detection of genomic regions associated with the disease. The GWAS results revealed DD incidence as a complex trait, where multiple genes control or regulate minor effects, with no specific genomic region highly contributing to the genetic variance. Although our sample size was limited, using gene set enrichment analysis (GSEA) significantly enriched gene ontologies associated with DD traits were identified. The significantly enriched GO terms from the ontology categories, biological process and cellular components, and molecular function, were highly involved in cellular and membrane function, revealing genes pertaining to adhesion, migration and proliferation. The processes are considered to be crucial for the pathogenesis of DD. This study provides a foundation for future studies in understanding the complex pathogenesis and genetic background of DD in beef cattle.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Compliance with ethical standards: The experimental procedures for the trial were approved by the Institutional Animal Care and Use Committee (IACUC V01525) at the University of Wisconsin-Madison, ensuring the ethical and sensitive care and use of animals in research, teaching and testing.
- Cramer G, Lissemore KD, Guard CL, Leslie KE, Kelton DF (2008) Herd- and Cow-Level Prevalence of Foot Lesions in Ontario Dairy Cattle. J Dairy Sci 91: 3888-3895. Link: https://bit.ly/3fJZ7b8
- Sullivan LE, Carter SD, Blowey R, Duncan JS, Grove-White D, et al. (2013) Digital dermatitis in beef cattle. Vet Rec 173: 582-582. Link: https://bit.ly/2AkNV4o
- Becker J, Steiner A, Kohler S, Koller-Bähler A, Wüthrich M, et al. (2014) Lameness and foot lesions in Swiss dairy cows: I. Prevalence. Schweiz Arch Tierheilkd 156: 71-78. Link: https://bit.ly/3dB63oT
- Krull AC, Shearer JK, Gorden PJ, Cooper VL, Phillips GJ, et al. (2014) Deep Sequencing Analysis Reveals Temporal Microbiota Changes Associated with Development of Bovine Digital Dermatitis. Infect Immun 82: 3359-3373. Link: https://bit.ly/35T0aRy
- Bruijnis MRN, Hogeveen H, Stassen EN (2010) Assessing economic consequences of foot disorders in dairy cattle using a dynamic stochastic simulation model. J Dairy Sci 93: 2419-2432. Link: https://bit.ly/2LiFuZp
- Bruijnis MRN, Beerda B, Hogeveen H, Stassen EN (2012) Assessing the welfare impact of foot disorders in dairy cattle by a modeling approach. Animal 6: 962-970. Link: https://bit.ly/2YSOE6X
- Read DH, Walker RL (1998) Papillomatous digital dermatitis (footwarts) in California dairy cattle: clinical and gross pathologic findings. J Vet Diagn Investi 10: 67-76. Link: https://bit.ly/2SWs1Lo
- Evans NJ, Brown JM, Scholey R, Murray RD, Birtles RJ, et al. (2014) Differential inflammatory responses of bovine foot skin fibroblasts and keratinocytes to digital dermatitis treponemes. Vet Immunol Immunopathol 16: 12-20. Link: https://bit.ly/2WlqwIu
- Holzhauer M, Bartels CJM, Döpfer D, van Schaik G (2008) Clinical course of digital dermatitis lesions in an endemically infected herd without preventive herd strategies. Vet J 177: 222-230. Link: https://bit.ly/3dD2vTg
- Scholey RA, Blowey RW, Murray RD, Smith RF, Cameron J, et al. (2012) Investigating host genetic factors in bovine digital dermatitis. Vet Rec 171: 624. Link: https://bit.ly/3fEv6cD
- Gomez A, Anklam KS, Cook NB, Rieman J, Dunbar KA, et al. (2014) Immune response against Treponema spp. and ELISA detection of digital dermatitis. J Dairy Sci 97: 4864-4875. Link: https://bit.ly/2WKGYkJ
- Schöpke K, Gomez A, Dunbar KA, Swalve HH, Döpfer D (2015) Investigating the genetic background of bovine digital dermatitis using improved definitions of clinical status. J Dairy Sci 98: 8164-8174. Link: https://bit.ly/3dOeP3d
- Biemans F, Bijma P, Boots NM, de Jong MCM (2017) Digital Dermatitis in dairy cattle: The contribution of different disease classes to transmission. Epidemics 23: 76-84. Link: https://bit.ly/2WOhaEk
- Kopke G, Waurich B, Wensch-Dorendorf M, Döpfer D, Rosner F, et al. (2017) Genetic parameters for improved phenotypes of susceptibility for digital dermatitis in Holstein dairy cattle. In: Proc. 19th Int Symp 11th Conf Lameness in Ruminants Munich, Germany. Link:
- Scholey RA, Evans NJ, Blowey RW, Massey JP, Murray RD, et al. (2013) Identifying host pathogenic pathways in bovine digital dermatitis by RNA-Seq analysis. Vet J 197: 699-706. Link: https://bit.ly/3crnAzH
- Refaai W, Ducatelle R, Geldhof P, Mihi B, El-shair M, et al. (2013) Digital dermatitis in cattle is associated with an excessive innate immune response triggered by the keratinocytes. BMC Vet Res 9: 193. Link: https://bit.ly/2Aiix6t
- van der Spek D, van Arendonk JAM, Bovenhuis H (2015) Genome-wide association study for claw disorders and trimming status in dairy cattle. J Dairy Sci 98: 1286-1295. Link: https://bit.ly/2WLApOS
- Peñagaricano F, Souza AH, Carvalho PD, Driver AM, Gambra R, et al. (2013) Effect of Maternal Methionine Supplementation on the Transcriptome of Bovine Preimplantation Embryos. PLoS ONE 8. Link:
- Abdalla EA, Peñagaricano F, Byrem TM, Weigel KA, Rosa GJM (2016) Genome-wide association mapping and pathway analysis of leukosis incidence in a US Holstein cattle population. Anim Genet 47: 395-407. Link: https://bit.ly/3bqq3t0
- Taye M, Kim J, Yoon SH, Lee W, Hanotte O, et al. (2017) Whole genome scan reveals the genetic signature of African Ankole cattle breed and potential for higher quality beef. BMC Genet 18: 11-11. Link: https://bit.ly/2SRfQzl
- Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4: 44-57. Link: https://bit.ly/3fCbdTD
- Kulow M, Merkatoris P, Anklam KS, Rieman J, Larson C, et al. (2017) Evaluation of the prevalence of digital dermatitis and the effects on performance in beef feedlot cattle under organic trace mineral supplementation1. J Anim Sci 95: 3435-3444. Link: https://bit.ly/3bpkRFI
- Döpfer D, Koopmans A, Meijer FA, Szakáll I, Schukken YH, et al. (1997) Histological and bacteriological evaluation of digital dermatitis in cattle, with special reference to spirochaetes and Campylobacter faecalis. Vet Rec 140: 620-623. Link: https://bit.ly/2zxFxhj
- Berry SL, Read DH, Famula TR, Mongini A, Döpfer D (2012) Long-term observations on the dynamics of bovine digital dermatitis lesions on a California dairy after topical treatment with lincomycin HCl. Vet J 193: 654-658. Link: https://bit.ly/3fFkw58
- Tremblay M, Bennett T, Döpfer D (2016) The DD Check App for prevention and control of digital dermatitis in dairy herds. Prev Vet Med 132: 1-13. Link: https://bit.ly/35Trr61
- Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, et al. (2007) PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am J Hum Genet 81: 559-575. Link: https://bit.ly/2X1qPrl
- Sargolzaei M, Chesnais JP, Schenkel FS (2014) A new approach for efficient genotype imputation using information from relatives. BMC Genomics 15: 478. Link: https://bit.ly/3fD8yc5
- Endelman JB (2011) Ridge Regression and Other Kernels for Genomic Selection with R Package rrBLUP. Plant Genome 4: 250-255. Link: https://bit.ly/2SX9tKI
- Varsos C, Patkos T, Oulas A, Pavloudi C, Gougousis A, et al. (2016) Optimized R functions for analysis of ecological community data using the R virtual laboratory (RvLab). Biodivers Data J 1: e8357. Link: https://bit.ly/3cq0edJ
- R Core Team (2017) Vienna, Austria: R Foundation for Statistical Computing. Link: http://bit.ly/2Mppho5
- SAS (2015) Cary, NC: SAS Institute Inc.
- Lippert C, Listgarten J, Liu Y, Kadie CM, Davidson RI, et al. (2011) FaST linear mixed models for genome-wide association studies. Nat Methods 8: 833-835. Link: https://go.nature.com/2WpeYEo
- Durinck S, Moreau Y, Kasprzyk A, Davis S, De Moor B, et al. (2005) BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21: 3439-3440. Link: https://bit.ly/2LmucDG
- Durinck S, Spellman PT, Birney E, Huber W (2009) Mapping Identifiers for the Integration of Genomic Datasets with the R/Bioconductor package biomaRt. Nat Protoc 4: 1184-1191. Link: https://bit.ly/3dEXAAW
- Zimin AV, Delcher AL, Florea L, Kelley DR, Schatz MC, et al. (2009) A whole-genome assembly of the domestic cow, Bos taurus. Genome Biol 10: R42. Link: https://bit.ly/3fFnG8O
- Yates A, Akanni W, Amode MR, Barrell D, Billis K, et al. Ensembl 2016. (2016) Nucleic Acids Res 44: D710-D716. Link: https://bit.ly/2AlHzSn
- Huang DW, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37: 1-13. Link: https://bit.ly/2xUSeTd
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, et al. (2000) Gene Ontology: tool for the unification of biology. Nat Genet 25: 25-29. Link: https://bit.ly/2YTPbFV
- The Gene Ontology Consortium (2015) Gene Ontology Consortium: going forward. Nucleic Acids Res 43: D1049-1056. Link: https://bit.ly/2WLCwlH
- Gene Ontology Consortium (2001) Creating the gene ontology resource: design and implementation. Genome Res 11: 1425-1433. Link: https://bit.ly/3cBpaiN
- Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, et al. (2009) AmiGO: online access to ontology and annotation data. Bioinformatics 25: 288-289. Link: https://bit.ly/3bv7NPi
- Holzhauer M, Hardenberg C, Bartels CJM, Frankena K (2006) Herd- and cow-level prevalence of digital dermatitis in the Netherlands and associated risk factors. J Dairy Sci 89: 580-588. Link: https://bit.ly/2LknkXh
- Rodriguez-Lainz A, Melendez-Retamal P, Hird DW, Read DH, Walker RL (1999) Farm- and host-level risk factors for papillomatous digital dermatitis in Chilean dairy cattle. Prev Vet Med 42: 87-97. Link: https://bit.ly/2WO5G3x
- Relun A, Lehebel A, Bruggink M, Bareille N, Guatteo R (2013) Estimation of the relative impact of treatment and herd management practices on prevention of digital dermatitis in French dairy herds. Prev Vet Med 110: 558-562. Link: https://bit.ly/2WmuGzZ
- Malchiodi F, Koeck A, Mason S, Christen AM, Kelton DF, et al. (2017) Genetic parameters for hoof health traits estimated with linear and threshold models using alternative cohorts. J Dairy Sci 100: 2828-2836. Link: https://bit.ly/2YU60R1
- Neupane M, Geary TW, Kiser JN, Burns GW, Hansen PJ, et al. (2017) Loci and pathways associated with uterine capacity for pregnancy and fertility in beef cattle. PLoS ONE 12. Link: https://bit.ly/3fJcwQr
- Friedl P, Mayor R (2017) Tuning Collective Cell Migration by Cell-Cell Junction Regulation. Cold Spring Harb Perspect Biol 9: a029199. Link: https://bit.ly/3cqKG9N
- Lawson CD, Burridge K (2014) The on-off relationship of Rho and Rac during integrin-mediated adhesion and cell migration. Small GTPases 5: e27958. Link: https://bit.ly/2LhvT5n
- Saeui CT, Mathew MP, Liu L, Urias E, Yarema KJ (2015) Cell Surface and Membrane Engineering: Emerging Technologies and Applications. J Funct Biomater 6: 454-485. Link: https://bit.ly/3dD4PJY
- Lee SE, Lee SH (2018) Skin Barrier and Calcium. Ann Dermatol 30: 265-275. Link: https://bit.ly/2LmMg0g
- Raguz J, Jeric I, Niault T, Nowacka JD, Kuzet SE, et al. (2016) Epidermal RAF prevents allergic skin disease. eLife 5: e14012. Link: https://bit.ly/2LmHEre
- Morselli E, Galluzzi L, Kepp O, Vicencio JM, Criollo A, et al. (2009) Anti- and pro-tumor functions of autophagy. Biochim Biophys Acta. 1793: 1524-1532. Link: https://bit.ly/2STDQ4D
- Vogel W, Gish GD, Alves F, Pawson T (1997) The Discoidin Domain Receptor Tyrosine Kinases Are Activated by Collagen. Mol Cell 1: 13-23. Link: https://bit.ly/3dD4Bm1
- Gumienny TL, Brugnera E, Tosello-Trampont AC, Kinchen JM, Haney LB, et al. (2001) CED-12/ELMO, a Novel Member of the CrkII/Dock180/Rac Pathway, Is Required for Phagocytosis and Cell Migration. Cell 107: 27-41. Link: https://bit.ly/3bnEaPL
- Sanders MA, Ampasala D, Basson MD (2009) DOCK5 and DOCK1 Regulate Caco-2 Intestinal Epithelial Cell Spreading and Migration on Collagen IV. J Biol Chem 284: 27-35. Link: https://bit.ly/2YVKKKQ
- Wang Y, Xu X, Pan M, Jin T (2016) ELMO1 Directly Interacts with Gβγ Subunit to Transduce GPCR Signaling to Rac1 Activation in Chemotaxis. J Cancer 7: 973-983. Link: https://bit.ly/3fMb00f
- Hernández-Vásquez MN, Adame-García SR, Hamoud N, Chidiac R, Reyes-Cruz G, et al. (2017) Cell adhesion controlled by adhesion G protein-coupled receptor GPR124/ADGRA2 is mediated by a protein complex comprising intersectins and Elmo-Dock. J Biol Chem 292: 12178-12191. Link: https://bit.ly/3cqwYUi
- Schreiber V, Moog-Lutz C, Régnier CH, Chenard MP, Boeuf H, et al. (1998) Lasp-1, a novel type of actin-binding protein accumulating in cell membrane extensions. Mol Med Camb Mass 4: 675-687. Link: https://bit.ly/2YSS21F
- Bai SW, Herrera-Abreu MT, Rohn JL, Racine V, Tajadura V, et al. (2011) Identification and characterization of a set of conserved and new regulators of cytoskeletal organization, cell morphology and migration. BMC Biol 9: 54. Link: https://bit.ly/2Af71c0
- Chen J, Den Z, Koch PJ (2008) Loss of desmocollin 3 in mice leads to epidermal blistering. J Cell Sci 121: 2844-2849. Link: https://bit.ly/3dGpIDN
- Vasilopoulos Y, Walters K, Cork MJ, Duff GW, Sagoo GS, et al. (2008) Association analysis of the skin barrier gene cystatin A at the PSORS5 locus in psoriatic patients: evidence for interaction between PSORS1 and PSORS5. Eur J Hum Genet 16: 1002-1009. Link: https://bit.ly/35VkiCy
- Zhang D, Zhu H, Harpaz N (2016) Overexpression of α1 chain of type XI collagen (COL11A1) aids in the diagnosis of invasive carcinoma in endoscopically removed malignant colorectal polyps. Pathol Res Pract 212: 545-548. Link: https://bit.ly/2YSuY34
- Tu H, Sasaki T, Snellman A, Göhring W, Pirilä P, et al. (2002) The Type XIII Collagen Ectodomain Is a 150-nm Rod and Capable of Binding to Fibronectin, Nidogen-2, Perlecan, and Heparin. J Biol Chem 277: 23092–23099. Link: https://bit.ly/3fJdUm7
- Stefanovic B, Stefanovic L, Schnabl B, Bataller R, Brenner DA (2004) TRAM2 Protein Interacts with Endoplasmic Reticulum Ca2+ Pump Serca2b and Is Necessary for Collagen Type I Synthesis. Mol Cell Biol 24: 1758-1768. Link: https://bit.ly/2zxHRVz
- Martin KR, Kantari-Mimoun C, Yin M, Pederzoli-Ribeil M, Angelot-Delettre F, et al. (2016) Proteinase 3 Is a Phosphatidylserine-binding Protein That Affects the Production and Function of Microvesicles. J Biol Chem 291: 10476-10489. Link: https://bit.ly/2LhwLXH
- Jensen MA, Wilkinson JE, Krainer AR (2014) Splicing factor SRSF6 promotes hyperplasia of sensitized skin. Nat Struct Mol Biol 21: 189-197. Link: https://bit.ly/2yRek9o
- Demaria M, Ohtani N, Youssef SA, Rodier F, Toussaint W, et al. (2014) An Essential Role for Senescent Cells in Optimal Wound Healing through Secretion of PDGF-AA. Dev Cell 31: 722-733. Link: https://bit.ly/2AkZ5pP
- Pastore S, Lulli D, Maurelli R, Dellambra E, De Luca C, et al. (2013) Resveratrol induces long-lasting IL-8 expression and peculiar EGFR activation/distribution in human keratinocytes: mechanisms and implications for skin administration. PloS One 8: e59632. Link: https://bit.ly/2LhFSHR
- Becker HM, Rullo J, Chen M, Ghazarian M, Bak S, et al. (190) α1β1 Integrin-Mediated Adhesion Inhibits Macrophage Exit from a Peripheral Inflammatory Lesion. J Immunol 190: 4305-4314. Link: https://bit.ly/35OzmSm
- Eble JA, Kassner A, Niland S, Mörgelin M, Grifka J, et al. (2006) Collagen XVI Harbors an Integrin α1β1 Recognition Site in Its C-terminal Domains. J Biol Chem 281: 25745-25756. Link: https://bit.ly/2SVD73a
- Agis H, Collins A, Taut AD, Jin Q, Kruger L, et al. (2014) Cell population kinetics of collagen scaffolds in ex vivo oral wound repair. PloS One 9: e112680. Link: https://bit.ly/3bqE2ig
- Lyle C, McCormick F (2010) Integrin αvβ5 is a primary receptor for adenovirus in CAR-negative cells. Virol J 7: 148. Link: https://bit.ly/2WnWsfl
- Zhang WM, Popova SN, Bergman C, Velling T, Gullberg MK, et al. (2002) Analysis of the human integrin α11 gene (ITGA11) and its promoter. Matrix Biol 21: 513-523. Link: https://bit.ly/3cqMfVd
- Zeltz C, Gullberg D (2016) The integrin–collagen connection – a glue for tissue repair? J Cell Sci 129: 653-664. Link: https://bit.ly/2WLFTcd
- Morandi EM, Verstappen R, Zwierzina ME, Geley S, Pierer G, et al. (2016) ITGAV and ITGA5 diversely regulate proliferation and adipogenic differentiation of human adipose derived stem cells. Sci Rep 6: 28889. Link: https://bit.ly/3dF07Ls
- Kabashima K, Sakata D, Nagamachi M, Miyachi Y, Inaba K, et al. (2003) Prostaglandin E2–EP4 signaling initiates skin immune responses by promoting migration and maturation of Langerhans cells. Nat Med 9: 744-749. Link: https://bit.ly/35OzGk2
- Denda S, Kumamoto J, Takei K, Tsutsumi M, Aoki H, et al. (2012) Ryanodine Receptors Are Expressed in Epidermal Keratinocytes and Associated with Keratinocyte Differentiation and Epidermal Permeability Barrier Homeostasis. J Invest Dermatol 132: 69-75. Link: https://bit.ly/2zxIHlb
- Denda M, Sokabe T, Fukumi-Tominaga T, Tominaga M (2007) Effects of Skin Surface Temperature on Epidermal Permeability Barrier Homeostasis. J Invest Dermatol 127: 654-659. Link: https://bit.ly/2AnRMhf
- Ehrenreiter K, Piazzolla D, Velamoor V, Sobczak I, Small JV, et al. (2005) Raf-1 regulates Rho signaling and cell migration. J Cell Biol 168: 955-964. Link: https://bit.ly/3bjEKhm
- Lock FE, Hotchin NA (2009) Distinct Roles for ROCK1 and ROCK2 in the Regulation of Keratinocyte Differentiation. PLOS ONE 4: e8190. Link: https://bit.ly/2WOw3pX
- Shen YJ, Kong ZL, Wan FN, Wang HK, Bian XJ, et al. (2014) Downregulation of DAB2IP results in cell proliferation and invasion and contributes to unfavorable outcomes in bladder cancer. Cancer Sci 105: 704-712. Link: https://bit.ly/2xSRFJp
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.


 Save to Mendeley
Save to Mendeley
